1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
|
reinitialize
import modevectors
fetch 1c3y, async=0
split_states 1c3y, 1, 1
split_states 1c3y, 23, 23
hide
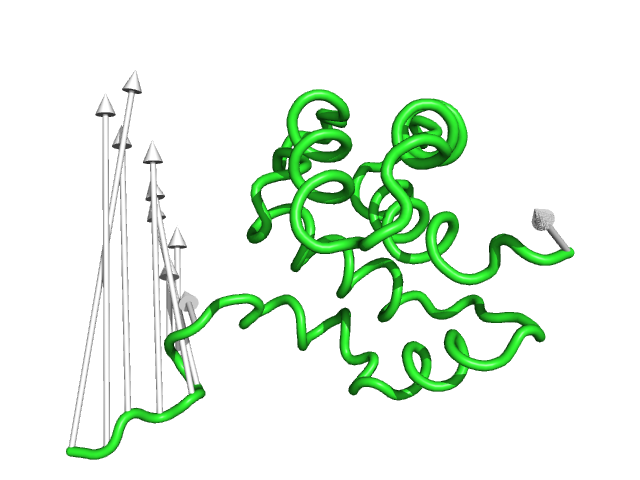
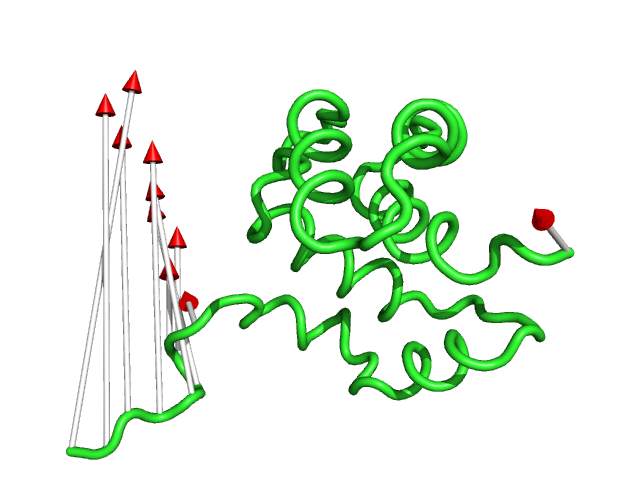
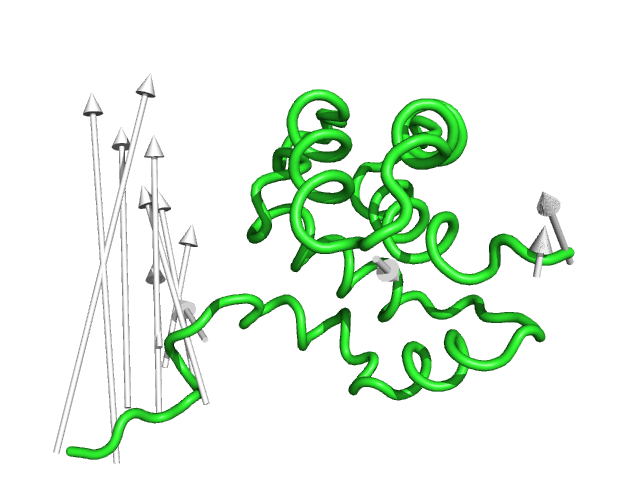
#This is the default setting (Fig.1)
modevectors 1c3y_0001, 1c3y_0023
#This is shows that the direction of the arrows is drawn from the 1c3y_0001 towards 1c3y_0023 (Fig.2)
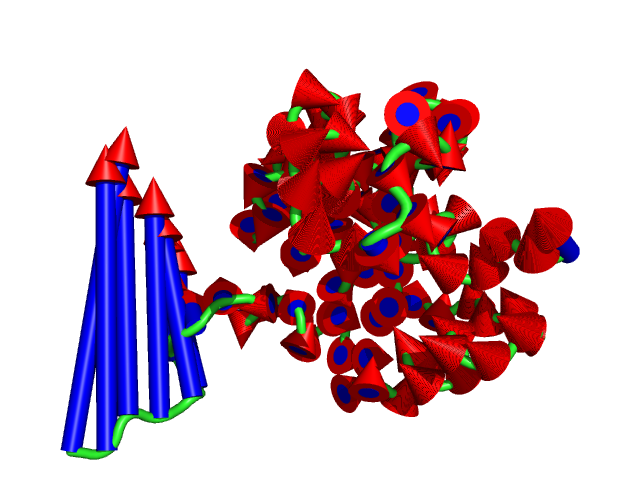
show cartoon, 1c3y_0023
color red, 1c3y_0023
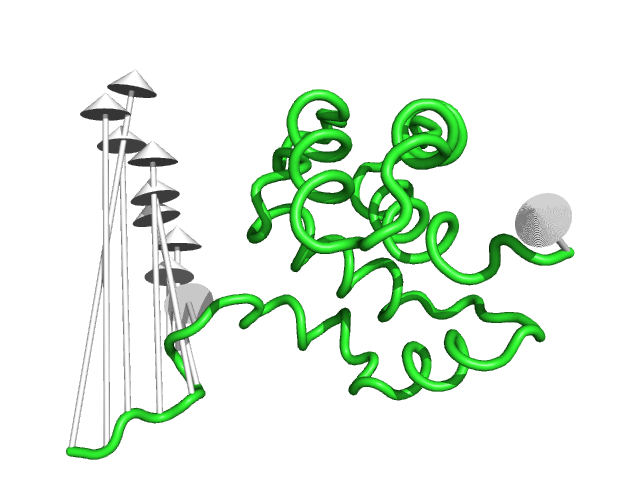
#This controls the base radius of the cone/arrow head in angstroms (Fig.3)
modevectors 1c3y_0001, 1c3y_0023, head=2.5
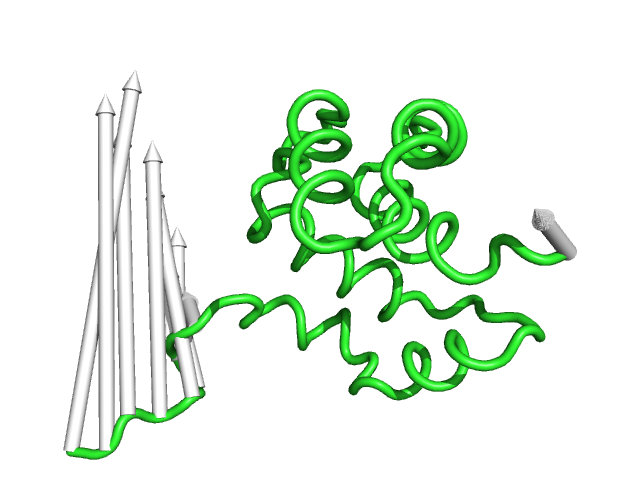
#This controls the radius of the cylinders/arrow tails in angstroms (Fig.4)
modevectors 1c3y_0001, 1c3y_0023, tail=0.75
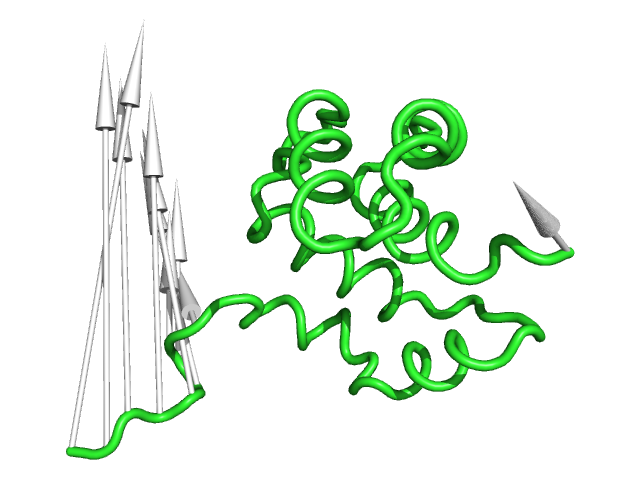
#This controls the length of the cone/arrow head in angstroms (Fig.5)
#Note that this option does NOT increase the vector length but simply changes the tail length
modevectors 1c3y_0001, 1c3y_0023, head_length=3.0
#This controls the colour of the cone/arrow head using RGB values (Fig.6)
modevectors 1c3y_0001, 1c3y_0023, headrgb=(1.0,0.0,0.0)
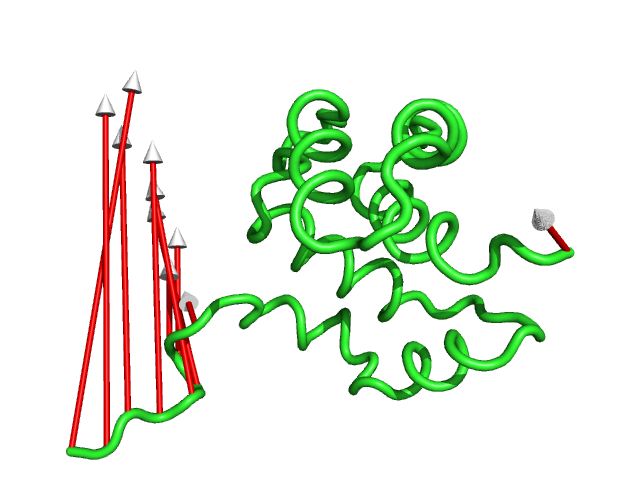
#This controls the colour of the cylinder/arrow tails using RGB values (Fig.7)
modevectors 1c3y_0001, 1c3y_0023, tailrgb=(1.0,0.0,0.0)
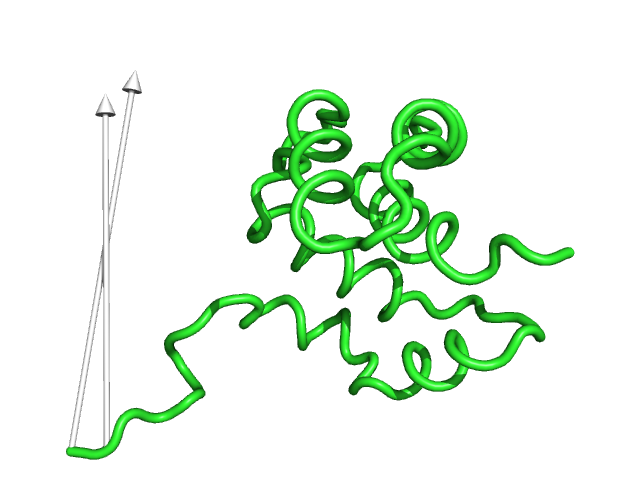
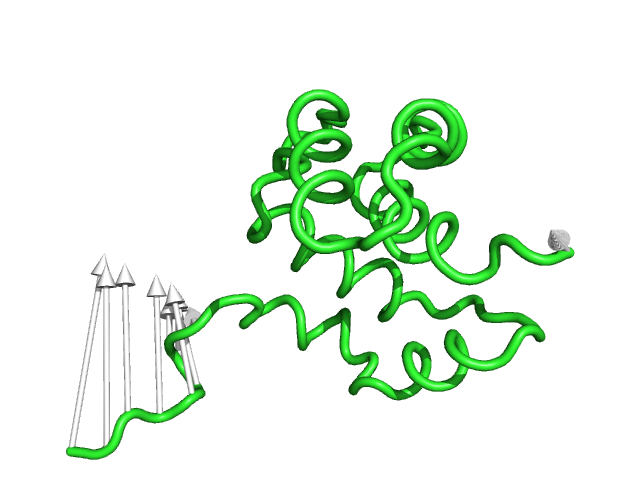
#This controls the which vectors to show based on a specific cutoff value in angstroms. Vector lengths that are less
#than the cutoff value will not be displayed (Fig.8)
modevectors 1c3y_0001, 1c3y_0023, cutoff=30.0
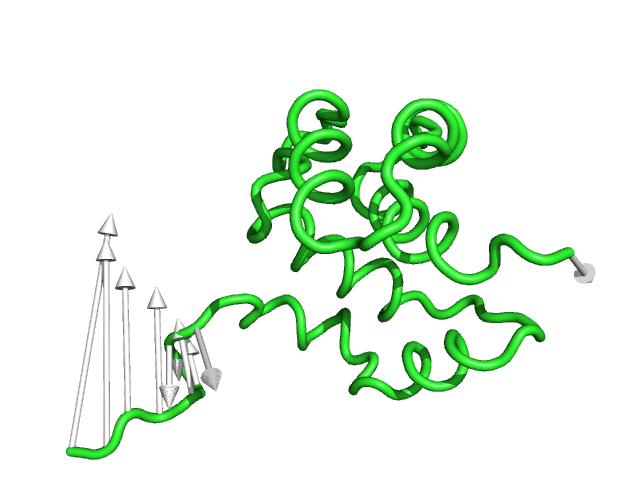
#This controls how many vectors to skip (integer value) and is useful when there are too many vectors showing.
#Skip=1 will show every other arrow (Fig.9)
modevectors 1c3y_0001, 1c3y_0023, skip=1
#This controls how much to cut from each vector (in angstroms). Note that arrows will point in the opposite directions
#when too much is cutoff (resulting in a negative vector length) (Fig.10) and should be used with caution!
modevectors 1c3y_0001, 1c3y_0023, cut=15.0
#This controls which atom to draw a vector from (Fig.11). Note that this is case-sensitive and is really only useful
#when atom=CA or when atom=P (for DNA backbones)
modevectors 1c3y_0001, 1c3y_0023, atom=CB
#This controls how much to multiple the length of each vector by (percentage increase/decrease) (Fig.12)
#This example halves the length of each vector (50%)
modevectors 1c3y_0001, 1c3y_0023, factor=0.5
#This hides the statistics which count atoms skipped, atoms counted (number of arrows showing), and number of atoms
#that did not meet the cutoff and are not shown
modevectors 1c3y_0001, 1c3y_0023, stat=hide
#This example shows multiple options being used (Fig.13)
modevectors 1c3y_0001, 1c3y_0023, head=2.0, tail=1.0, head_length=2.0, headrgb=(1.0,0.0,0.0), tailrgb=(0.0,0.0,1.0),
cutoff=0.0,skip=0,cut=0,atom=CA,factor=0.8
#Finally, this example hides all arrow tails and only uses arrow heads via the notail option(No Figure)
modevectors 1c3y_0001, 1c3y_0023, head=2.0, cutoff=0.0,skip=0,cut=0,atom=CA,factor=0.8, notail=1
|